The rapid spread of the COVID-19 virus highlights the need for an open analytics environment so that the global research community can collaborate on developing effective countermeasures.
(27 February, Hinxton, UK) Open access data and open, reproducible analytics are crucial for fast and efficient response to global health crises and can significantly contribute to our understanding of novel viruses. This is the main point of a joint paper by Galaxy teams from the United States, Australia and ELIXIR who re-analysed available COVID-19 data and assessed the reproducibility of initial papers on the COVID-19 genome.
The authors highlight the lack of primary data for the COVID-19 virus and call for greater transparency in order to effectively respond to public health emergencies: “Infectious disease outbreaks often occur in locations where infrastructure necessary for data analysis may be inaccessible (...) there is a global need to ensure access to free, open, and robust analytical approaches that can be used by anyone in the world to analyze, interpret, and share data.”
The authors’ assessment of the availability of the data on COVID-19 reveals that public access to sequence data that were used to assemble the COVID-19 genome is lacking. This can significantly slow down the development of efficient treatments since sequence data can be used to uncover viral diversity.
“Only one of the four papers that presented the COVID-19 genome provided access to the raw data. Without the raw data, the entire analysis presented in the three manuscripts is completely unverifiable and irreproducible,” says Björn Grüning, one of the authors of the paper and co-Lead of the ELIXIR Tools Platform. “This is simply unacceptable. We now have the tools and infrastructure which enable any researcher to make their analytical procedures 100% reproducible and transparent,” adds Grüning.
Supporting rapid, reproducible research during global health crises
To demonstrate the maturity of the public infrastructure and community-curated software for biomedical data analysis, the authors re-analysed all available raw COVID-19 data. Their results showed that the analyses described in the original papers presenting the COVID-19 genome can be reproduced on public infrastructure and using open source tools.
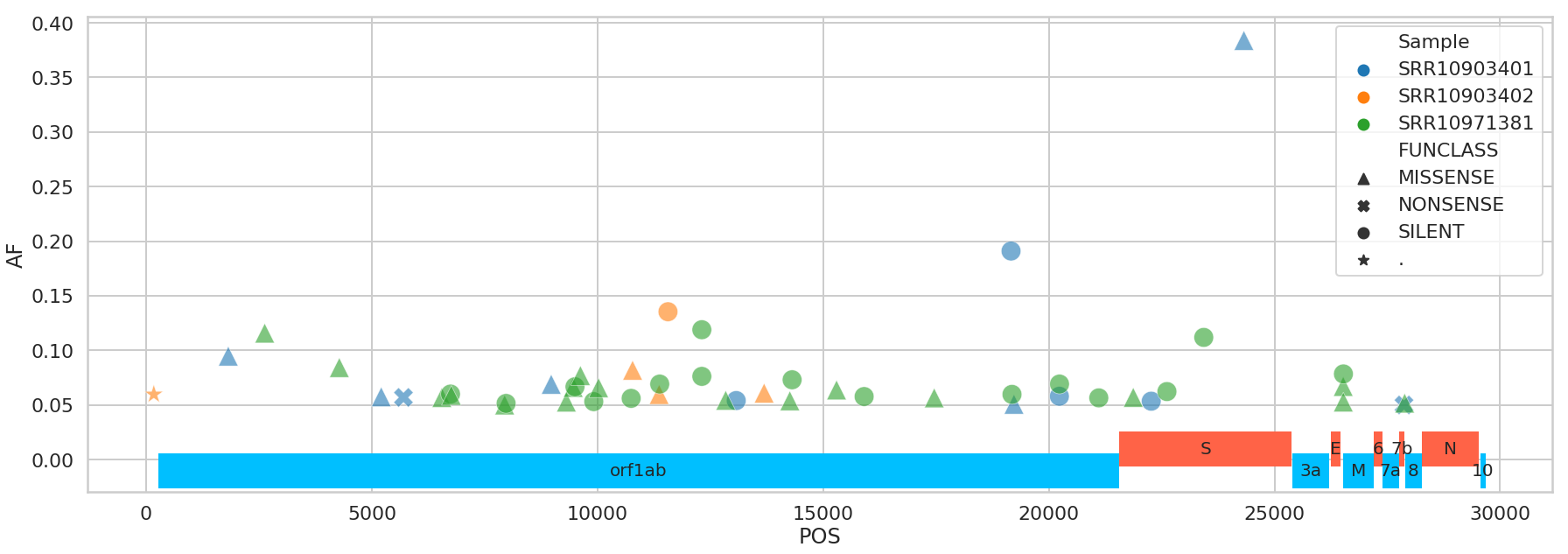
The presented analyses were performed using only free software and deployed on four Galaxy platforms: in the USA and Australia, and in two platforms operated by ELIXIR Nodes in Germany (Galaxy Europe) and Belgium (Galaxy Belgium).
Thanks to this effort, researchers worldwide now have access to the data, the complete analysis pipeline and the computational power necessary for their own analyses of the COVID-19 data. This means that newly published data can be re-analysed within hours and compared with existing data.
Global research market of ideas
ELIXIR is working towards enabling researchers to freely access and reuse data, tools, analysis workflows and computational power to derive novel insights to develop an effective response to global threats to public health. ELIXIR’s continuous support to Galaxy and other community-based infrastructure projects plays a crucial role in developing such infrastructure.
A necessary condition in this development is a concerted effort in developing and delivering bioinformatics training to researchers at every stage in their careers. Reaching the critical mass of the biomedical research community in disseminating best practices in data analysis is essential in improving accessibility and reproducibility of biomedical research.
At the same time, by engaging with funders and publishers we need to ensure that researchers are incentivised to publish their data and data analysis pipelines.
The authors conclude their paper with a call to policymakers to open up the “global research market”, where competition arises from deriving understanding rather than exclusive access to samples and data.
“Other disciplines have embraced the benefits of global data generation and sharing, astronomy and high energy physics being two highly successful examples. We have the opportunity to mirror their successes in infrastructure funding by demonstrating that biological research can embrace the same global perspective on common infrastructure investment and data sharing.”
Read the full paper:
No more business as usual: agile and effective responses to emerging pathogen threats require open data and open analytics
usegalaxy.org, usegalaxy.eu, usegalaxy.org.au, usegalaxy.be and hyphy.org development teams
bioRxiv 2020.02.21.959973; doi: https://doi.org/10.1101/2020.02.21.959973
All analyses presented in the paper are fully documented and accessible at:
See also:
- Coronavirus data analysis: News release by the Galaxy Europe team, Albert-Ludwigs-Universität Freiburg
- Galaxy Australia contributes to global research effort into COVID-19, News release by The Australian BioCommons
- UseGalaxy, SARS-CoV-2 / COVID-19, and how effective responses to emerging pathogen threats ..., News release by the Galaxy project